The advent of Long Axial Field-of-View (LAFOV) PET scanners has shifted the dosimetry paradigm in PET/CT
imaging. The high sensitivity of these systems allows for substantial reductions in radiotracer activity, rendering
the volumetric CT component the dominant source of ionizing radiation. For dose-sensitive populations such as
pediatric and obstetric cohorts, eliminating the volumetric CT entirely is highly desirable. However, the CT serves
a dual purpose: providing anatomical context and enabling attenuation correction (AC) for PET reconstruction,
as the attenuation map is typically derived directly from the CT. Similarly, whole-body studies acquired on
PET/MRI systems require estimation of the attenuation map from MR images. In both scenarios, the absence of
a CT poses a reconstruction challenge.
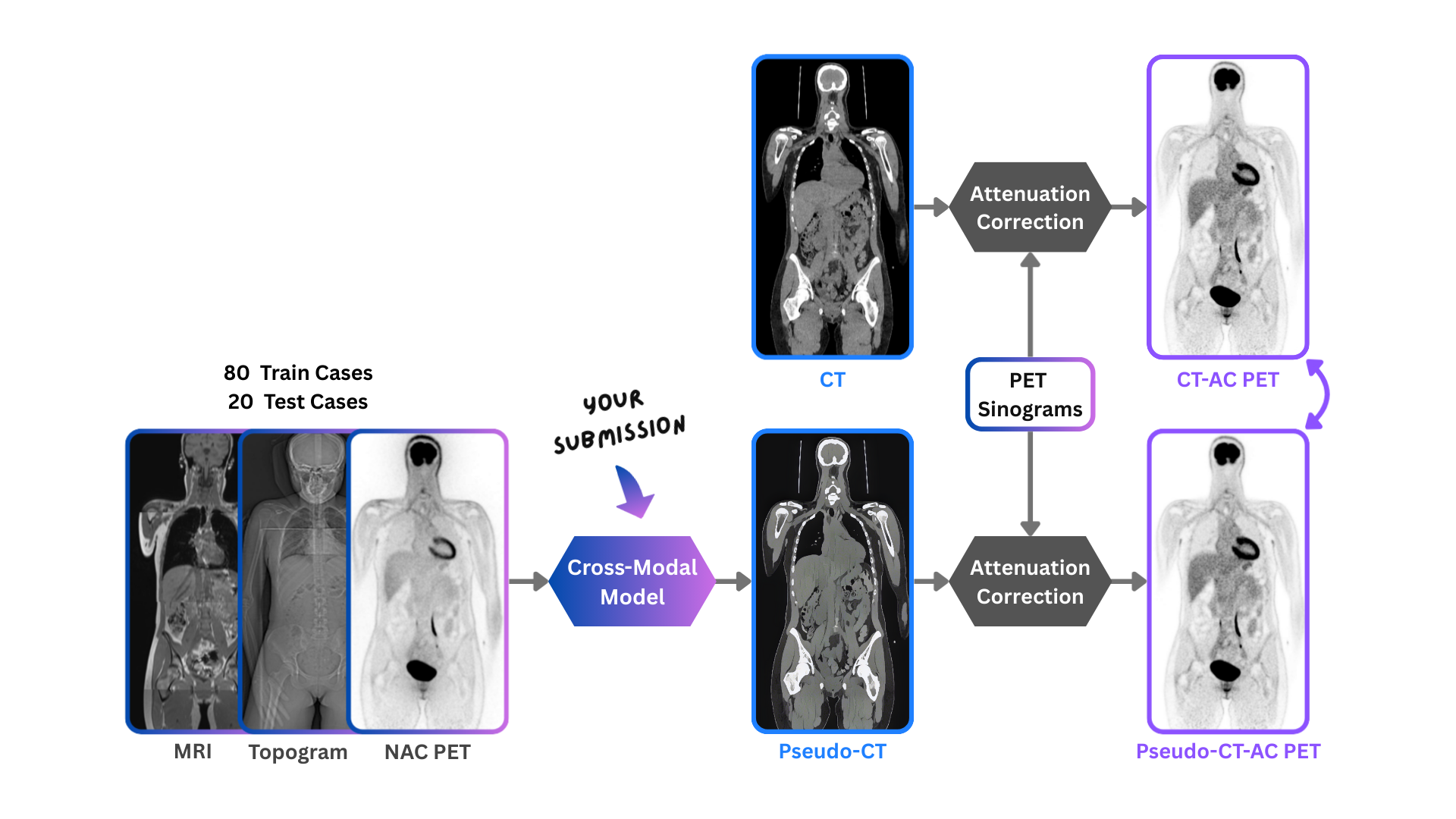
To address this, the Big Cross-Modal Attenuation Correction (BIC-MAC) challenge tasks participants with
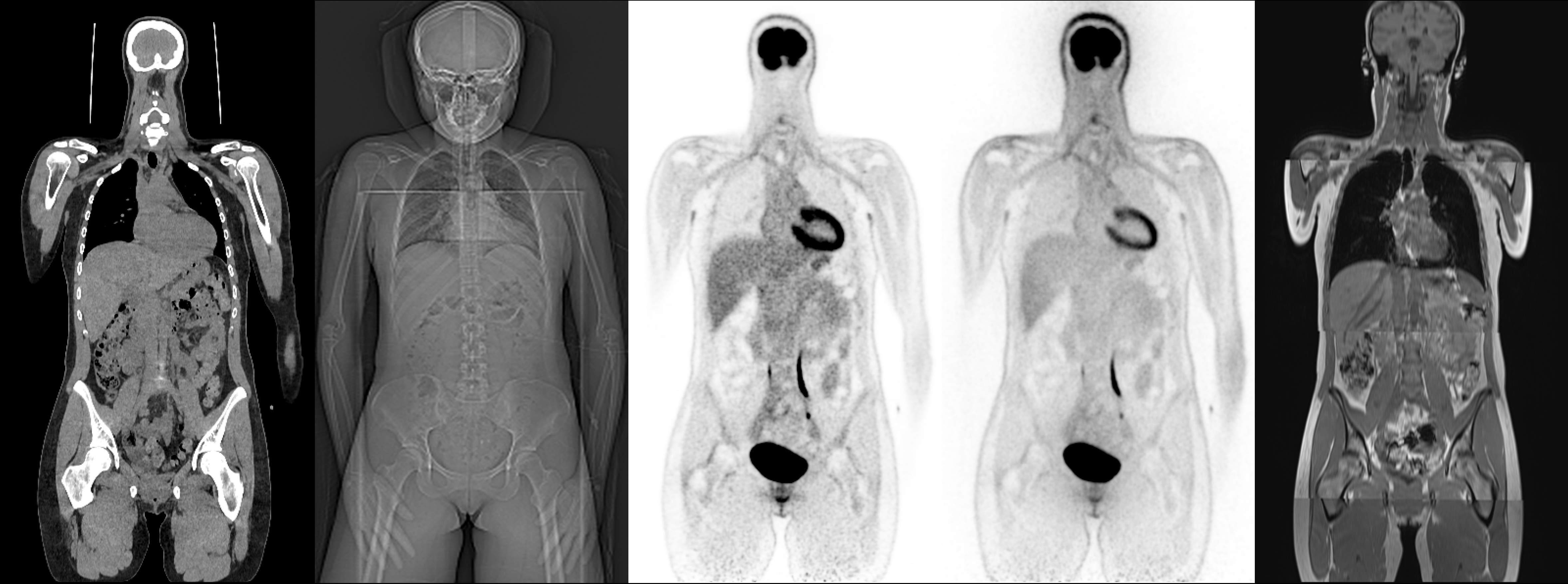
synthesizing a 3D pseudo-CT from other available modalities. We present a novel multimodal dataset comprising
whole-body PET, CT, Topogram (scout radiograph), and MRI for 100 healthy volunteers. The cohort is age- and
sex-stratified, with data acquired on Siemens Biograph Vision Quadra and Siemens MAGNETOM Vida
scanners. Participants will receive a training set of 75 cases containing Non-Attenuation Corrected (NAC)
[18F]FDG PET images, scan-planning Topograms, and same-day DIXON MRI, alongside reference CT and CT-based attenuation-corrected PET (CTAC-PET) images. Critically, we also provide scatter maps, sinograms, and
Docker containers with open-source reconstruction software, enabling closed-loop optimization on the training
set - a capability previously restricted to hospital sites with access to proprietary vendor software.
The challenge comprises a single task: generate a pseudo-CT from the available input modalities. The pseudo-CT will be used to reconstruct PET images, which are then quantitatively compared against reference CTAC-PET images. Both static and dynamic PET reconstructions are evaluated to assess downstream accuracy across
different clinical contexts. A defining technical characteristic of this challenge is the integration of modalities with
different dimensionalities and acquisition geometries. While the 3D NAC-PET and 2D Topograms are spatially
aligned with the target attenuation map, both lack anatomical detail. In contrast, whole-body MRI offers high
bone and soft-tissue contrast but is acquired in a different scanner geometry with different patient positioning
and body deformations. Consequently, participants must develop algorithms capable of fusing spatially
unaligned information from 3D volumetric MRI with that of the 3D NAC-PET and 2D Topograms.